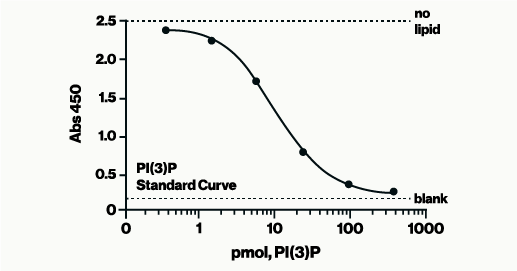
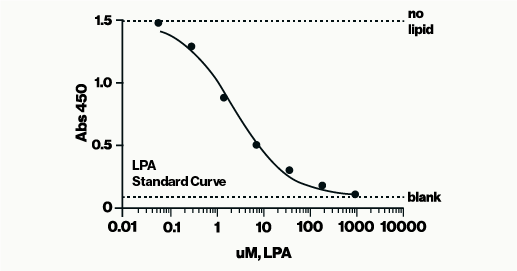
The PI(3)P Mass ELISA measures the amount of PI(3)P extracted from cells by means of a competitive ELISA.
Assay Range: 400 to 0.39 pmol PI(3)P
Sample Type: Dried down lipid extracted from cells
Sample Volume: 180 µL/sample, run in triplicate.
Product Background
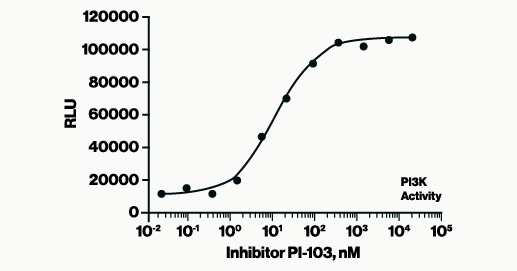
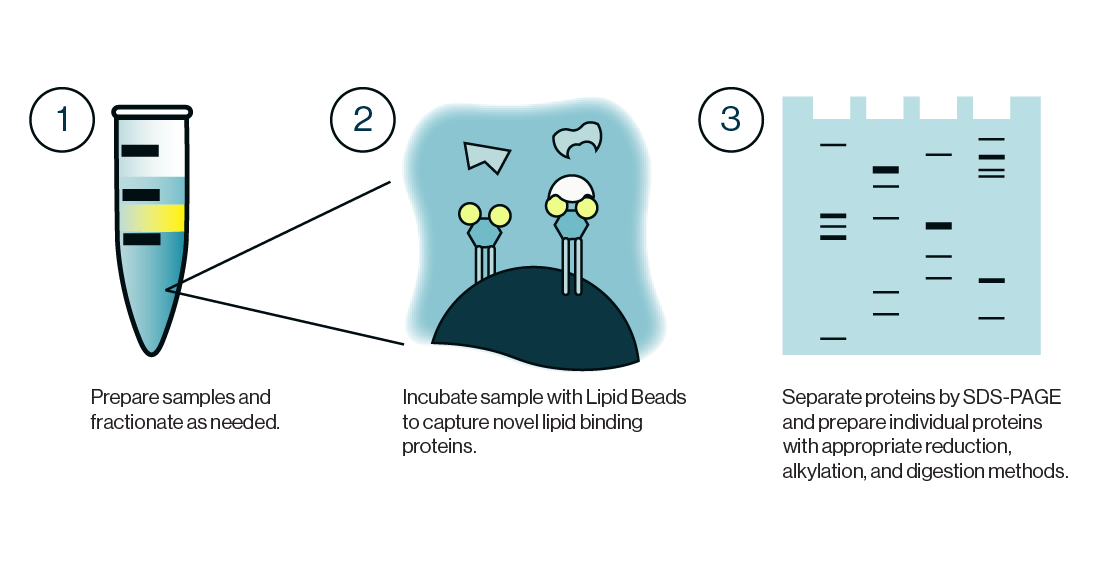
The PI(3)P Mass ELISA Kit, a 96-well competitive ELISA, allows for the determination of PI3-K activity by quantifying the amount of PI(3)P found in cells. After following a non-radioactive lipid extraction protocol, the PI(3)P samples obtained from millions of cells are detected on a competitive ELISA. The cellular PI(3)P samples are mixed and incubated with a PI(3)P Detector protein, then added to a PI(3)P-coated microplate for competitive binding. A peroxidase-linked secondary detector and colorimetric detection is used to detect the PI(3)P protein binding to the plate. The assay is sensitive to 1.0 pmol PI(3)P. The colorimetric signal is inversely proportional to the amount of PI(3)P obtained.
Keywords: PI3K, PI3 Kinase, PI3 K, PI3-K, PI3P